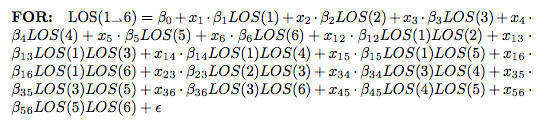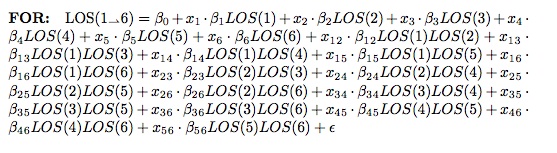There is a developing data science, machine learning, and deep learning community in the South Florida area that I support. The topics were diverse, from business intelligence to online ad buying to health tech. I was invited by Data Science Salon to attend and I was really pleased to do so.
The conference was hosted by Formulated.by and was held in Miami’s CIC near University of Miami/Jackson Memorial Hospital. It was a two day conference – I attended only the second day.
Vendors participating and hosting the conference were: Dataiku, Vertica, Plot.ly, and Formulated.ly, O’Reilly, Alteryx, & Domino Data Lab.
Here is the Thursday conference agenda:
I got through the traffic in Miami just in time to make the tail end of the Meditation exercise. I’ll be honest – talking about data science gets me excited, so I really wasn’t in the mood to calm down. Miami traffic also doesn’t make me calm down. But it was fun, nonetheless.
Brian MacDonald of the Florida Panthers started off with an interesting presentation about how the Florida Panthers, as an organization, solved the problem of how much to charge for their seats in a game, which ended up being a very traditional data science problem, beginning with an exploration of the data, discerning relationships in that data, and then creating predictive models. It turns out that the demand of seats is related to: day of week, opposing team, home team performance, holidays (some were highly negative, like Valentine’s day) and how late in the season the game is being played. They utilized a regression model controlling for independent variables, and thereafter were able to predictively model sales, attendance, and even season ticket holder renewals.
Michael Conway from Bidtellect spoke on their self-service predictive analytics platform for online ad bidding – which is using the Vertica service. Eye opening (for me as a physician) that they are participating in 15,000,000,000 (yes, number is accurate) auctions daily for online ad placement. Communicated that engagement rates are important, and by measuring post-click consumer activity you can document the value of the ad.
The data science team of Kevin U and Mark Fridson from Carnival Cruise Lines spoke – this was a really excellent talk, first about digital transformation of a traditional Fortune 500 company, and then some nuts & bolts. Kevin hammered home the importance of having a data-driven culture, which flows from the highest levels of the organization to spur adoption and deal with “change management” (That exists in healthcare too, by the way). One reality of being in South Florida was the skills gap – qualified data people are hard to come by.
Mark discussed the importance of multichannel engagement via snail mail, email, and social media as digital channels – sharing insights closely tied to generational cohorts. For each age group, Carnival has an “ideal customer” which they try to match as closely as possible to Boomers respond best via snail mail (USPS), while Gen X and Millenials use email and social media. For Generation Z, its all social media, but for different purposes. Snapchat creates exposure, while Instagram represents captured moments. Facebook is for acquaintance update and communication, and Twitter is most useful for interest and influencers. I thought that breakdown was particularly useful for those in marketing.
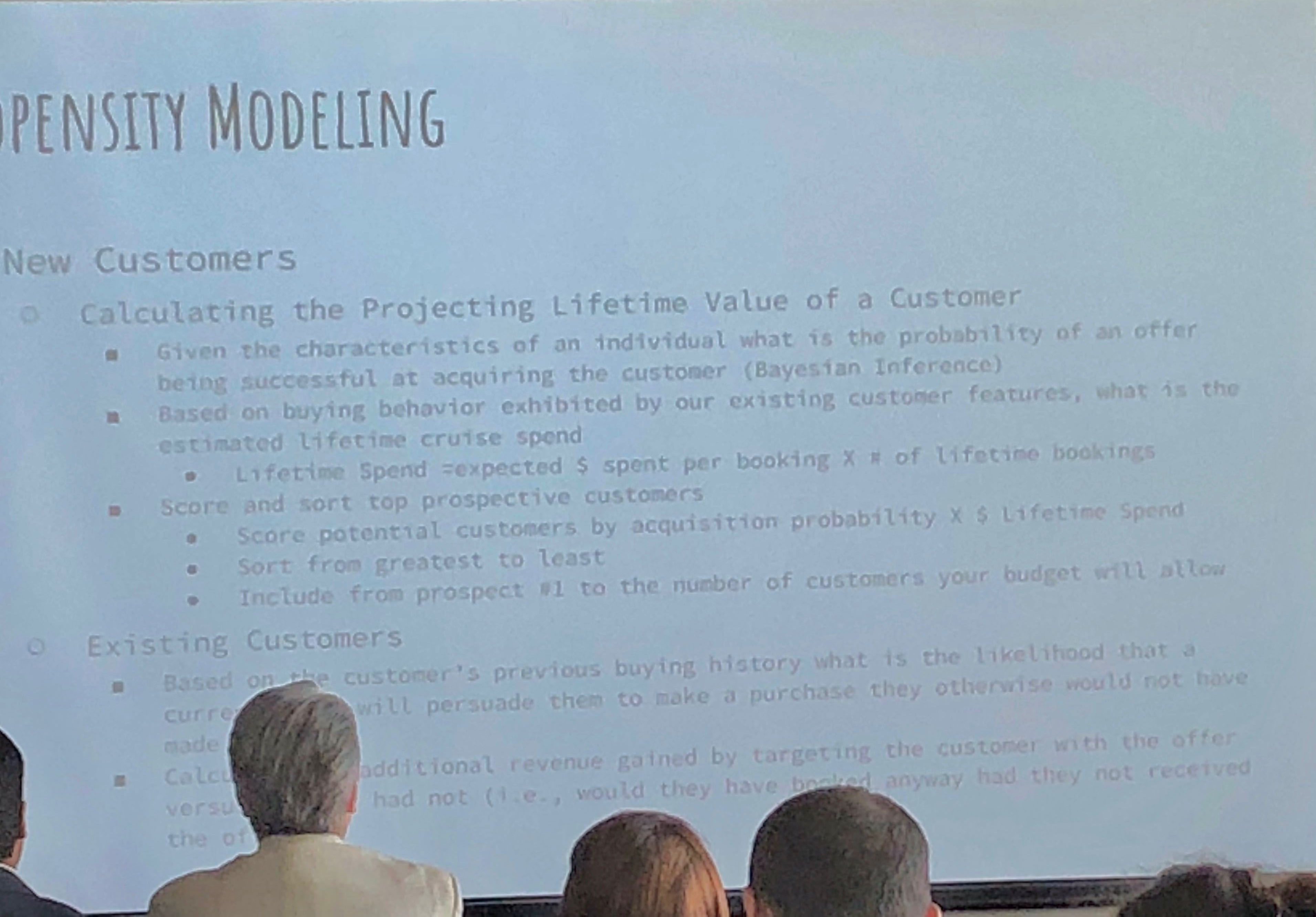

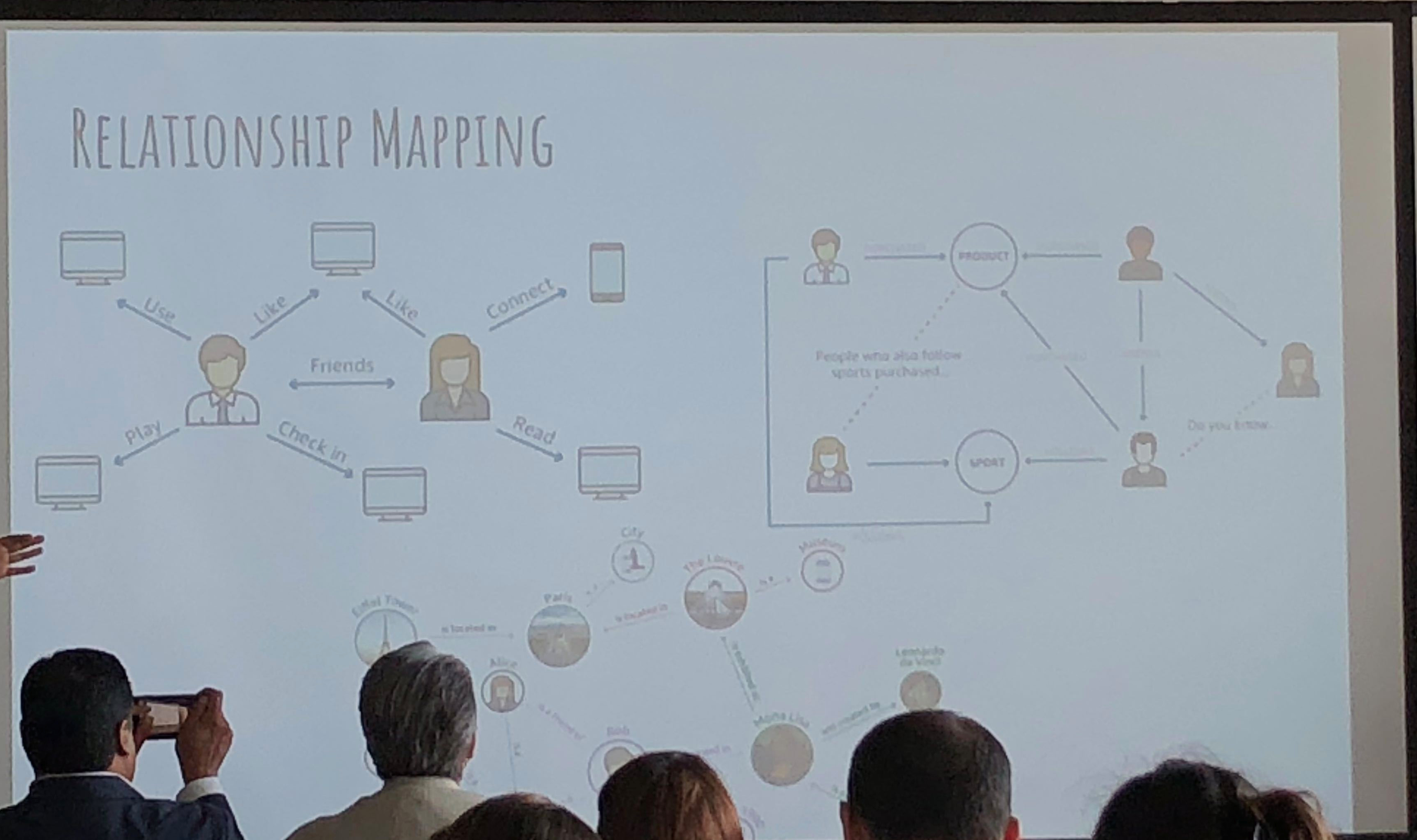
They use propensity modeling to calculate CLV (Customer Lifetime Value) and use Bayesian analysis. Content personalization is performed with: demographics, frequency, booking patterns, after-purchase add-ons, and even an element of serendipty (! – remember that piece on antifragility I did? These guys get it). They do use social relationship mapping and have been using some NLP text analysis but feel its hard to use AI NLP in SoMe.
Catalina Arango next spoke, and her talk was non-technical, aimed at beginners and managers desiring to implement data science elements in their enterprises. I took this opportunity to speak to the Dataiku and Vertica folks as this was a refresher for me.

Next up was Alex Rubynstein from Mt. Sinai in NYC – Mt . Sinai is One of the more proactive medical centers in the country regarding analytics and recognizing the value of data. I have seen them advertising for multiple positions to monetize their research.
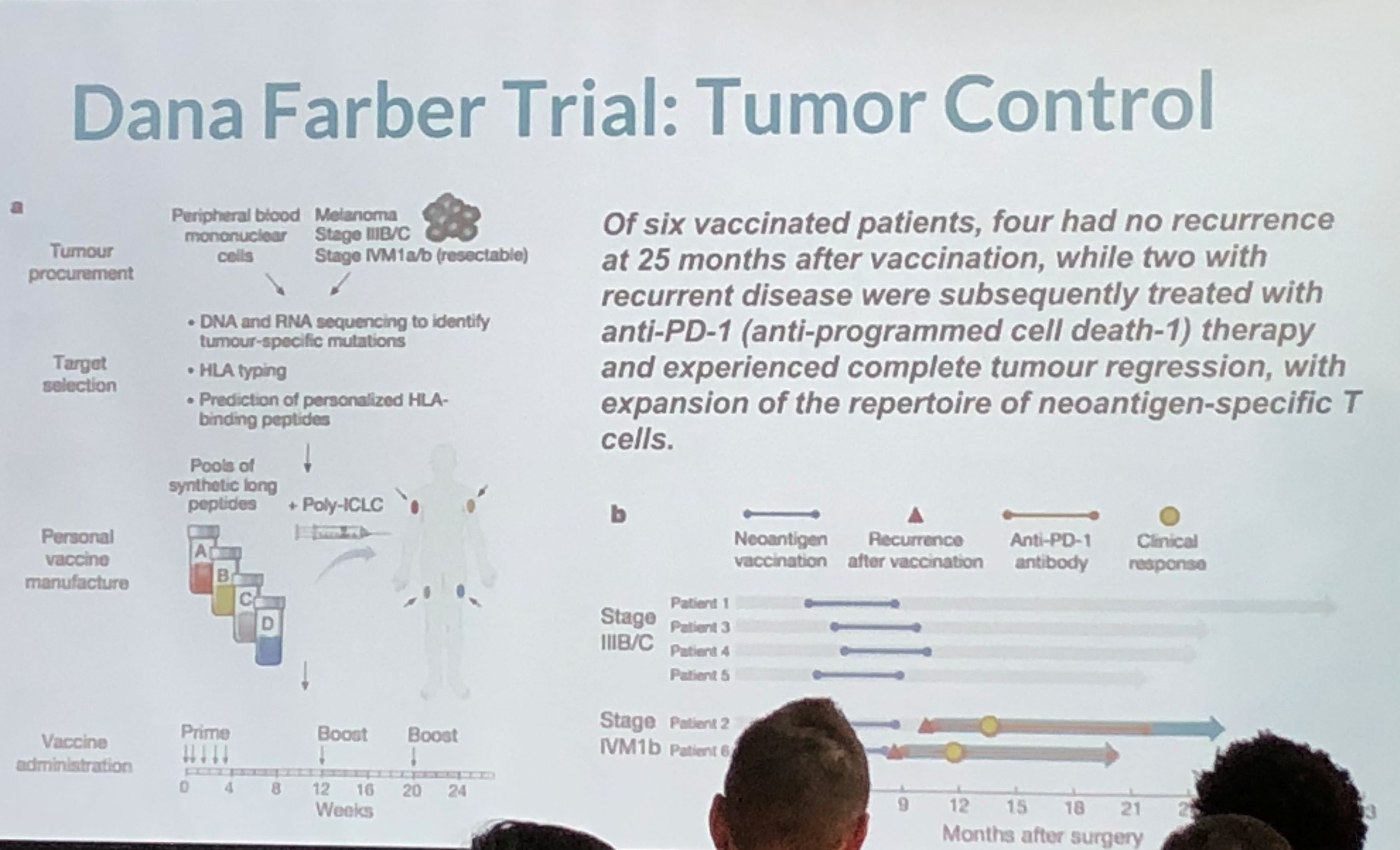
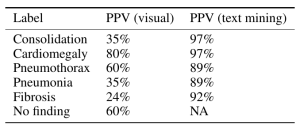
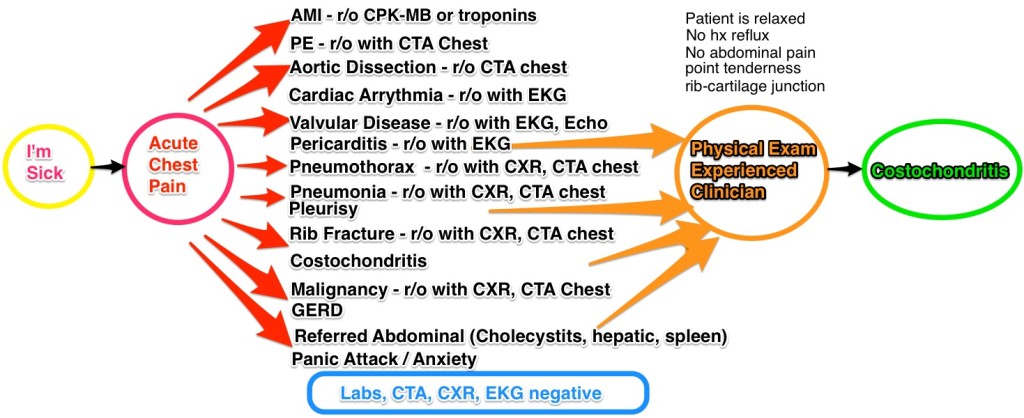
This was an interesting take on personalized medicine and genomics using big data for analysis. Because of cancer’s lethality, more experimentation is possible which have resulted in some novel therapies which approach cure, or at least transforming cancer into a chronic condition. The cancer vaccine approach treats the patient’s immune system to either enhance the immune response (to overcome immune suppression) or to increase the sensitivity of the immune system to the cancer (to overcome immune escape). They take the patient’s gene sequence, and the tumor gene sequence, filter the two and target on the order of 5-20 mutations, combining the vaccine with an adjuvant. They use machine learning on the candidate targets as the number of mutations exceeds the number of targets. They are continuing to expand on their sample size, which is extremely small, and because of the individualized nature of the therapy, very costly. Nevertheless, early results are promising. The primary limitation is the individualized and handcrafted nature of the vaccine.
Lunch followed – Subway sandwich boxes, which were fine. Networking at a data science conference can be tough (stereotypes anyone?) but I managed to find a few good folks to chat with.
A panel followed composed of three speakers – Dr. Irma Fernandez, chief academic officer of St. Thomas University; Colleen Farrelly, Data Scientist at Kaplan; Mauro Damo, chief data scientist at Dell; and Anton Antonov, Consultant at Accendo Data. There were a broad number of topics discussed. Main points were the following: Publishing data can be damaging, so be aware of what you are putting out there. Narrow AI only at this time – no general AI (we know that)! This was a good, in-the-fields survey of current trends and issues.

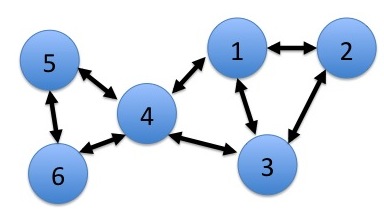
Athanassios Kintaskis, Sr. Machine Learning engineer at Capital One had an interesting presentation on MCL (Markov Clustering) Sparse Graphs – this was a good technical talk, some of which went over my head. As opposed to K-means clustering algorithms which are sensitive, but can’t tell you how many groups are present (you need to choose), this approach simulates random walks in a graph and uses a flow dynamic to create clusters.
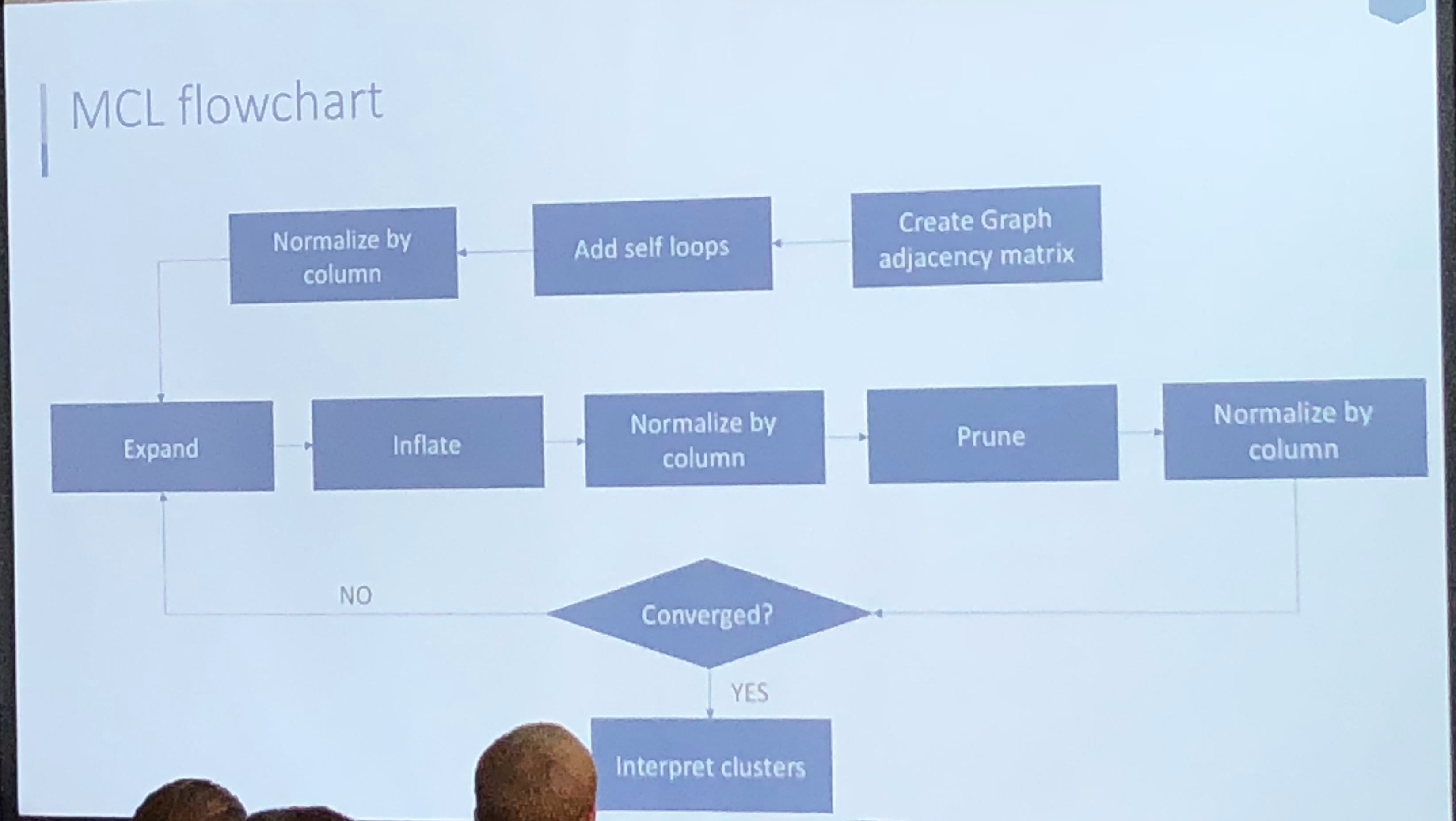

Markov Chain transitions can be modeled as a matrix, and that’s about as far as I got before I was interrupted by a phone call. This was an interesting and meaty talk, and I probably need to read up more on the topic before publically displaying my ignorance.
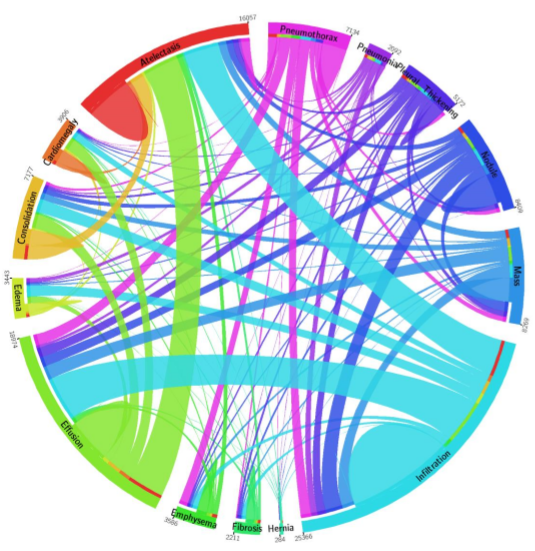
Anabetsy Rivero of Metastatic AI gave a nice introductory presentation on Convolutional Networks in medical imaging (head over to my other blog: www.ai-imaging.org for more on that or read my prior articles on this). Anabetsy is a machine learner that is focusing on breast cancer diagnostics.
There were a few other presentations but this is a blog, not a manifesto!
All in all, I appreciated what Formulated.by did to bring this type of conference to Miami. It is a necessary part of growing the Miami Data Science community, and I would love to see more events like Data Science Salon in the future. A 2nd DataScienceSalon:Miami is slated for November 6-7, 2018.

FULL DISCLOSURE: Because of my involvement in the South Florida Data Science and Machine Learning community, I received complimentary entrance.