Note: Update 4/22/20: Since this post was first written there has been a great deal of information released about the novel coronavirus COVID19, with an equal amount of confusion. Many preprints have been released to satisfy the hunger for information about this new and concerning disease. Unfortunately, there are both problems in quality and veracity and source material must be considered, as not all is of the same quality, and frankly, some might be intentional misinformation.
After this blogpost was published, Radiology leadership from the organized specialty societies came out with the recommendation NOT to use imaging on COVID-19 patients for diagnosis, but only for investigation of a worsening clinical course not explainable by COVID. Many AI projects were initiated of extremely poor/questionable quality, using as few as 5 cases for training. (!) I’ll instead suggest an adaptation of the 2018 RSNA pneumonia challenge winner which is being hosted on the Arterys platform if you are looking for such a product . (Full Disclosure: I have no financial interest in Arterys or the makers of that product)
I’ve tried hard to avoid hype for the purposes of this blog. So as to the question – is AI useful in detecting COVID via imaging? At this time I don’t know – further imaging data and study will be needed. My hope is that the point is somewhat moot, as if accurate rapid PCR tests and antibody testing are widely available, the need for detection via imaging will be less of a concern, and I think that is why the specialty societies argued against imaging as a front line diagnostic tool.
First post in a while. This was supposed to be “Wither Value? – an update on Value Based Care in 2020” But that can wait as the coronavirus situation in China gives us pause.
First off, our hearts go out to the afflicted in China, especially Wuhan. Diseases like COVID afflict randomly and pass sentence indiscriminately upon their victims, uncaring of race, creed, national origin, or political affiliation. We can all appreciate the primary risk is simply being alive and in the wrong place at the wrong time. As human beings, we all are potentially vulnerable.
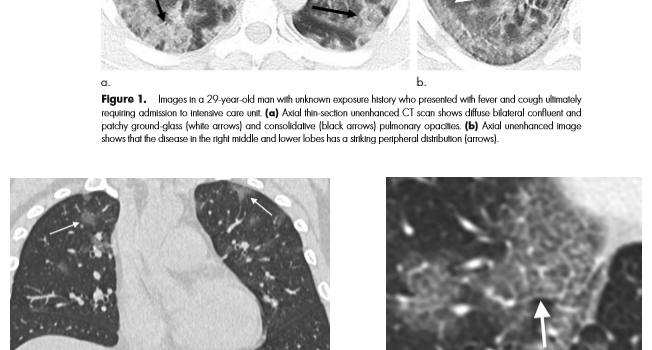
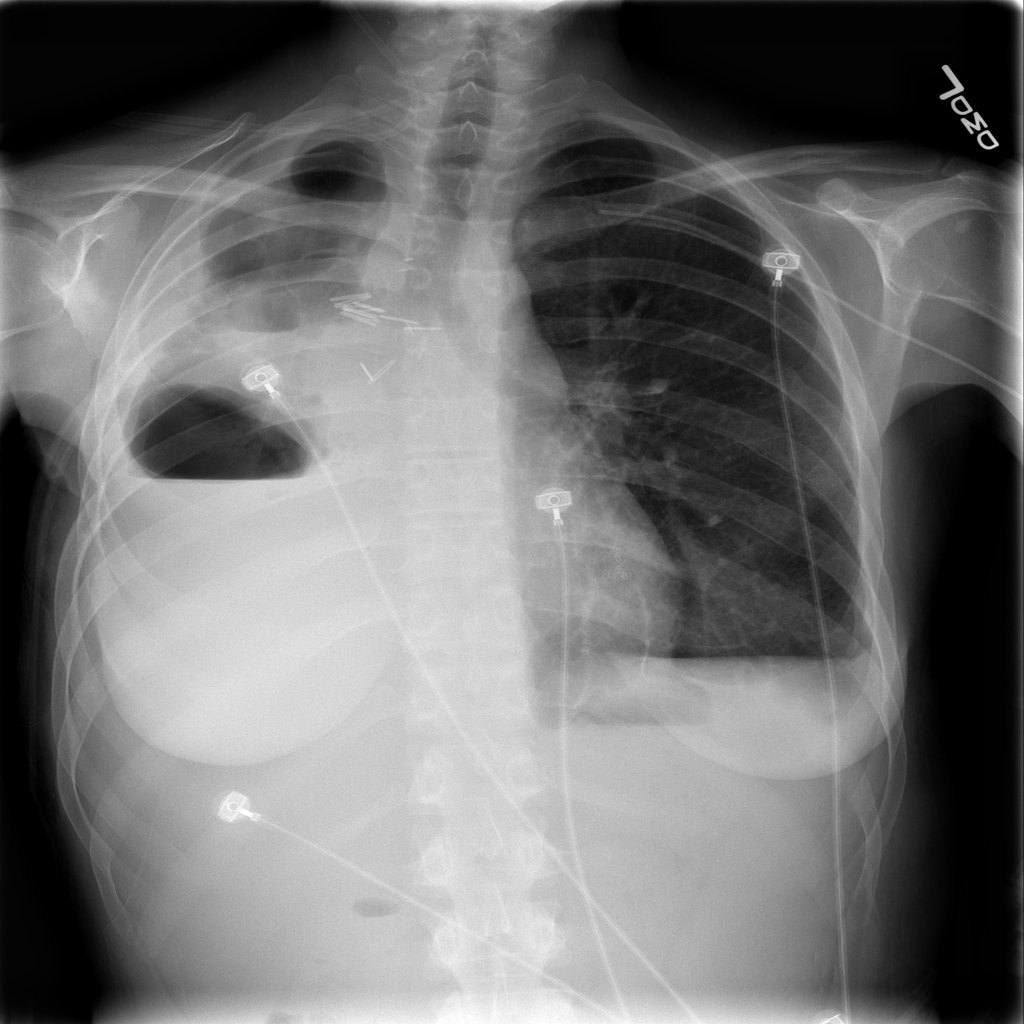
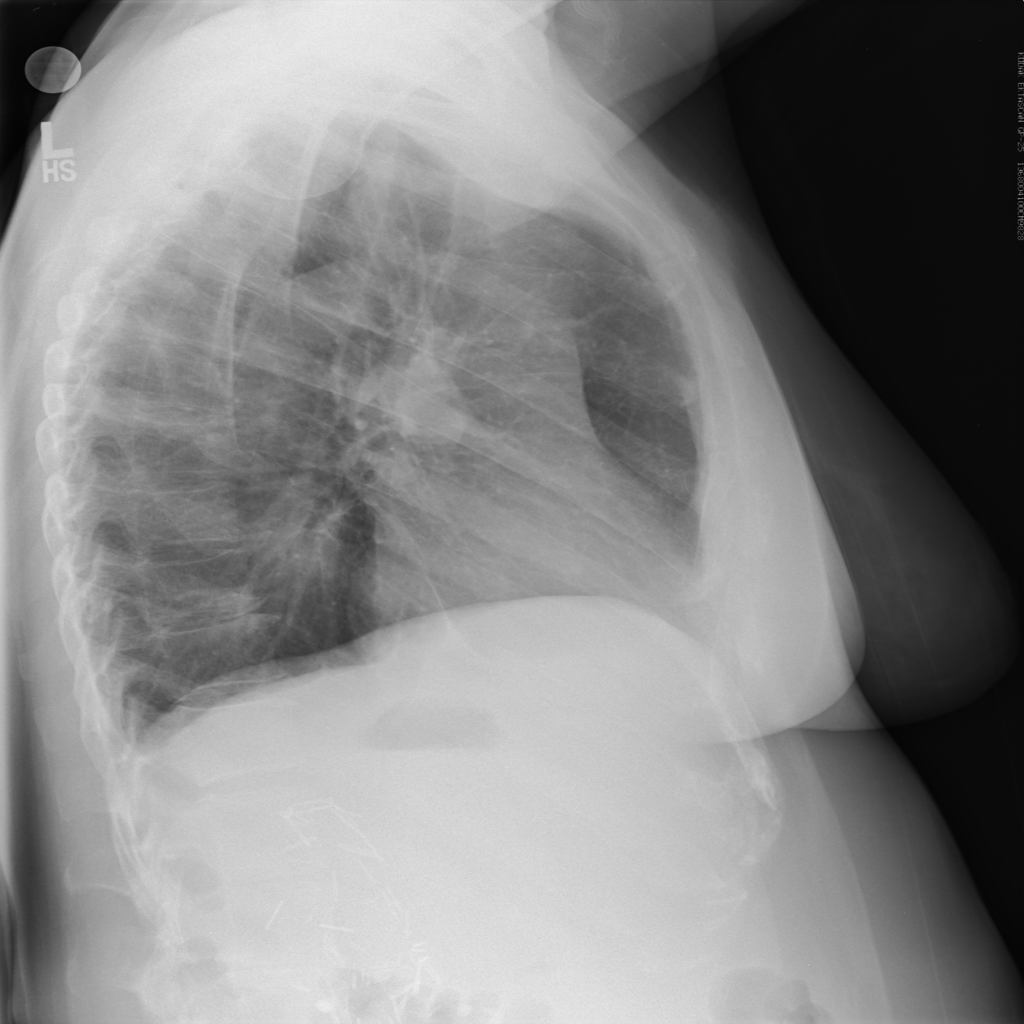
I write this after the nearly doubling of reported cases following a change in diagnostic criteria, first brought to my attention 2/12/20 by Scott Gottlieb, former FDA commissioner. Previously, the only reported cases were those confirmed by PCR, and some people commented that cases were rising steadily at about 3000/day. That might have been the capacity limit of national PCR testing. Clearly though, imaging (specifically a CT scan showing bilateral patchy ground glass opacities which are fairly typical of viral pneumonitis but nonspecific) is becoming part of the diagnostic algorithm, and revealing a more extensive distribution which was suspected from anecdotal social media reports.
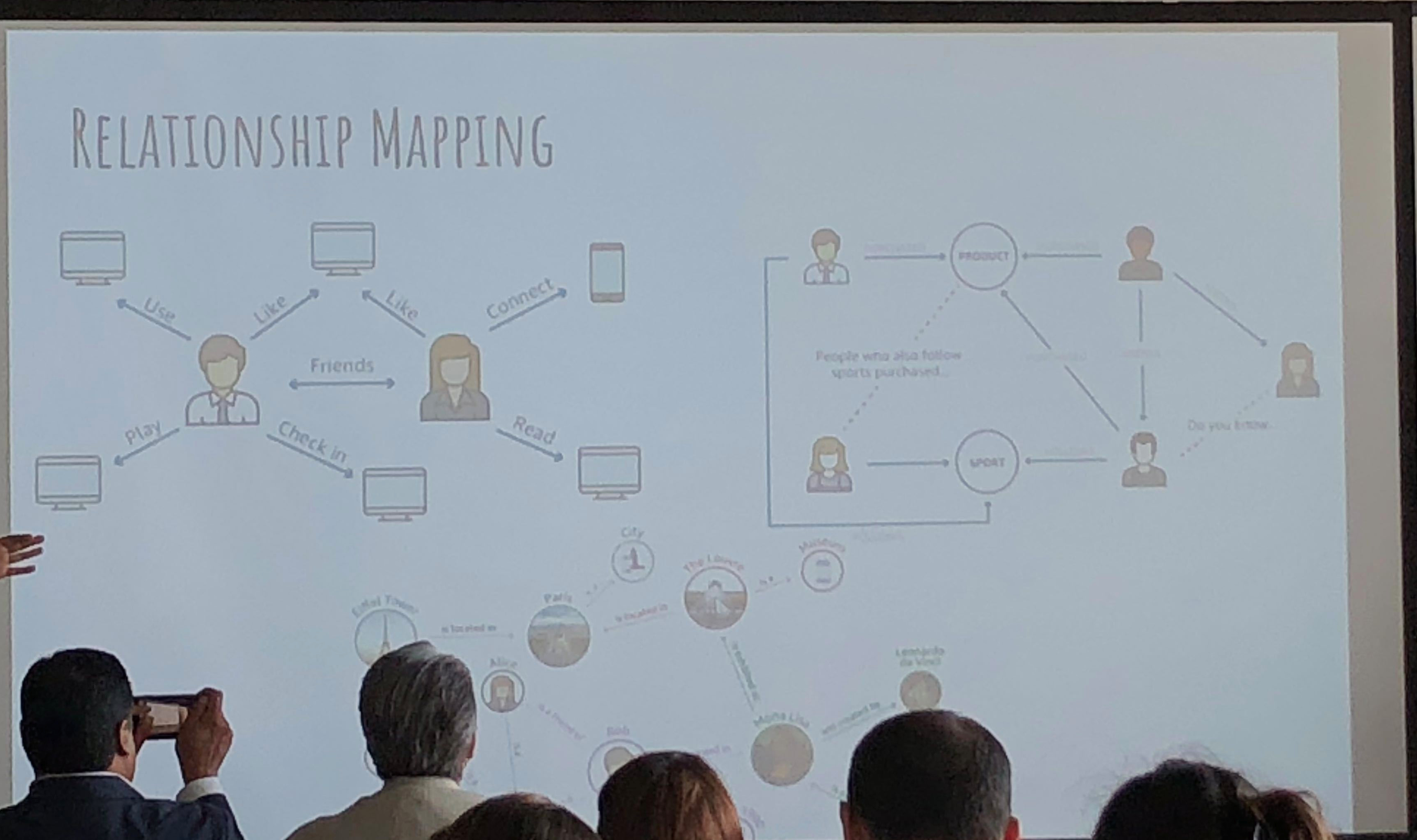
Which begs the question – could AI help?
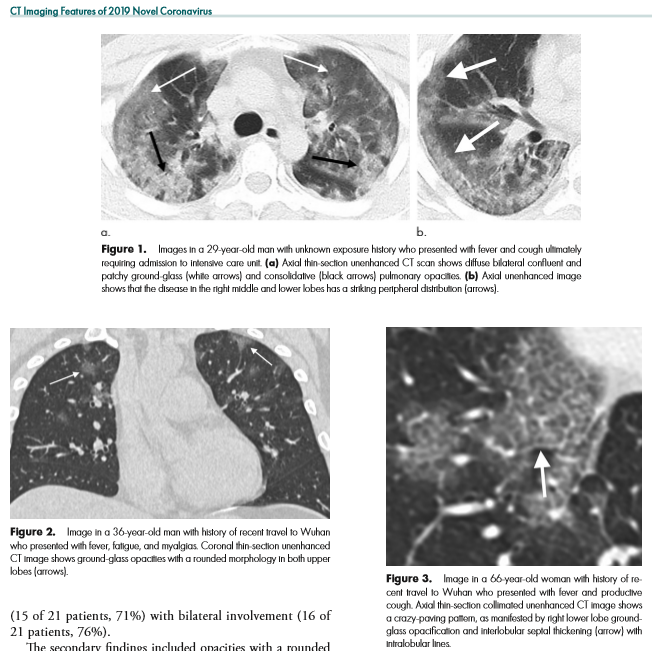
As CT imaging now is part of the diagnostic criteria, we have an opportunity to take positive CT scans from patients with positive COVID-19 PCR and make an initial, standardized dataset which could be in DICOM format. It should be as high quality as possible, and ideally have the ground glass opacities and progressive consolidation / ARDS-like appearance labeled with bounding boxes. If that Chinese startup with $100 million in funding doesn’t have the appropriate tools, my friend George over at MD.AI does. Once the basic dataset is created, a second dataset with similar, presumptive cases based upon the new WHO criteria (CT scan +, Wuhan contact, lymphopenia) can be assembled with weak labeling, perhaps aided by some semi-supervised learning techniques to make the labeling less onerous (similar to what we did in this paper). In order not to make it an ARDS detector, initially presenting imaging should be used. A similar number of controlled negative cases should be assembled for the dataset with a negative label. Then a classifier can be trained for detection / absence of disease.
While such a CT-based classifier would prove helpful in the developed world and urban China, the developing world is not helped by this CT classifier. For that reason, it would be prudent to create a dataset of standard chest radiographs from the ground truth positive patients identified in the two CT datasets above. This would hold value, with or without the bounding boxes and could be used for screening purposes.
Chest X-ray based deep learning screening would be useful for the rest of the world where not only access to advanced imaging is limited, but also access to radiologists. Studies can wait for up to 60 days to be interpreted by a physician in some developing countries. Chest X-ray is also less bulky and more portable, and could also be used at borders and immigration control points in conjunction with temperature checks to assess for potential geographic spread of COVID-19.
Part of the Value of AI in this circumstance is the rapid reproduction and dissemination of knowledge in front of a clear and present danger. Physicians might simply not know of COVID-19’s findings or not recognize them because they are too unlikely in their practice. Therefore, those first few cases slip by, and allow for increased local infections. The ability to be proactive is valuable. The dataset, and to a lesser extent any actual model represents a rapid transfer of knowledge among physicians. Dissemination of the dataset, and eventual AI classifier is limited only by internet access and end-user integration. The cost of developing such a single-purpose classifier would be miniscule – on the order of a research grant. More is being lost by economic slowdown on an hourly basis.
With the dataset open sourced, new cases could be added either on a collective basis via a registry created by governmental or NGO’s for this purpose or administered by a respective country’s health department/ministry. Additional cases could be added, and supervised by local academic physicians, who would be best suited to detect spectrum shift in disease presentation if the virus should mutate and change its pathologic presentation.
So, this post is predominantly aimed at my Asian colleagues who are simply by virtue of their location, present at the epicenter of this illness. To the healthcare professionals of the middle kingdom – it must be difficult to be witness to what you are seeing – I cannot even personally conceive it. You have a long tradition of excellence in the healing arts, stretching back to Hua Tuo of the Three Kingdoms and beyond. Receive my heartfelt respect for taking up the rod of Aesculapius at personal risk to yourself in order to minister to the needs of others. In conjunction with your close research colleagues, you are in a unique position to create this database – you have the clinical case knowledge and the PCR data. Furloughed or idle technologists, residents, and radiology physicians can annotate the data you provide, not only providing +disease / -disease labels on CT slice images, but bounding boxes on the areas of pulmonary involvement. While you are doing that, take the presenting chest x-ray and do the same! Those steps, by the way, can be done from home, over the internet. Once assembled into a high quality labelled dataset of a sufficient number of cases, release it publicly for the good of all people and nations. Models can be created and shared on the internet once the data is released. At the last RSNA I saw a neat open-source data lake for imaging data /DICOM called KHEOPS – perhaps this would be a way to accomplish this across users and platforms.
I’m aware that this post might not age well. On the other hand, I have a bit of a bully pulpit so I am going to use it. It is far too easy to fall either into hysteria or indifference when faced with a fearsome circumstance. I believe that rationality combined with empathy is the best response.