Author: Stephen Borstelmann MD
Opportunistic detection of type 2 diabetes using deep learning from frontal chest radiographs
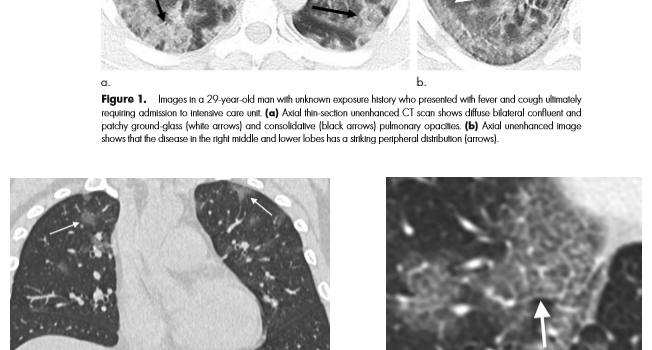
Combining AI and Value – the COVID 19 coronavirus as a potential use case.
Do we need more medical imaging?

Data Science Salon: Miami
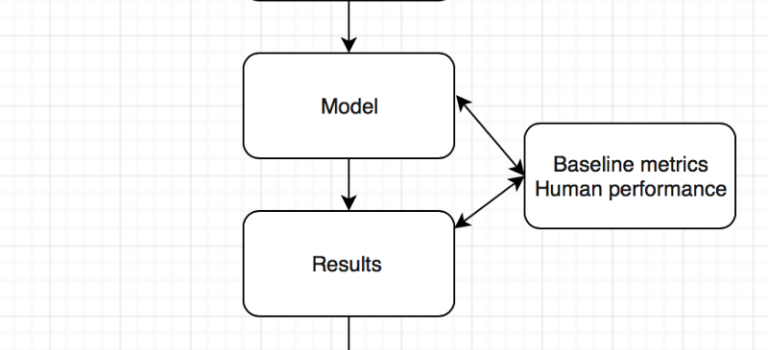
Are computers better than doctors ? Will the computer see you now ? What we learnt from the ChexNet paper for pneumonia diagnosis …

OODA loop revisited – medical errors, heuristics, and AI.
